Note
Go to the end to download the full example code or to run this example in your browser via Binder.
Handling OOSM using inverse time dynamics
In previous examples, we have shown how to deal with out-of-sequence measurements (OOSM) by using a fixed time-delay, or a time buffer. This example focuses on a different approach to deal with OOSM.
As shown in literature (e.g., [1]) there are multiple approaches on how to deal with OOS measurements. Simply ignoring them, assuming that the fraction of these is small and it will not, significantly impact the quality of the tracking; iterating over last \(\ell\) measurements with a fixed lag (see what it is called Algorithm A and B in [2] and [3], and the other OOSM examples) or, like in this example, you can include any OOSM and re-process the measurement using inverse-time dynamics creating pseudo-measurements.
This approach is known in literature as Algorithm C (from [2], [3]) and it will be referred to as such later on.
To explain how it works, we consider a single target scenario. This algorithm deals with the presence of delayed measurements by using the predicted dynamics of the target to go back in time from the actual arrival time and obtain a pseudo-measurement obtained from simulating the detection at the “expected” scan time. In this manner we can reconstruct the measurements chain without discarding any information, and we don’t need to store a large fix-lag distribution of data (as can happen with other algorithms).
In this example, we focus our efforts in showing how to use the algorithm C in a multi-target scenario with clutter. We simulate two sensors obtaining scans of two objects travelling with a nearly constant velocity transition model and with some level of clutter. At specific timesteps we insert a delay in one of the scans from the sensor and we employ Algorithm C to process in the correct way the chain of measurements.
To visualise the benefit of the algorithm, we also include a tracker which ignores OOSM completely. We also employ track-to-truth metrics to highlight the differences.
This example follows this structure:
Create the ground truths scans and detections of the targets;
Instantiate the tracking components;
Run the tracker and apply the algorithm on delayed measurements;
Run the comparison between the resulting tracks and plot the results.
General imports
from datetime import datetime, timedelta
import numpy as np
from copy import deepcopy
# Simulation parameters
start_time = datetime.now().replace(microsecond=0)
np.random.seed(2000) # fix a random seed
num_steps = 65
scans_delay = 5 # seconds between each scan
extra_delay = 2 # external delay in some scans
delay = 0 # empty delay
prob_detect = 0.90 # Probability of detection
lambdaV = 2 # Parameters for the clutter
v_bounds = np.array([[-1, 450], [-130, 130]]) # Parameters for the clutter
clutter_spatial_density = lambdaV/(np.prod(v_bounds[:, 0] - v_bounds[:, 1])) # Clutter spatial density
Stone Soup imports
from stonesoup.models.transition.linear import CombinedLinearGaussianTransitionModel, \
ConstantVelocity
from stonesoup.types.groundtruth import GroundTruthPath, GroundTruthState
from stonesoup.types.state import GaussianState, State, StateVector
from stonesoup.types.update import GaussianStateUpdate # needed for the track
1. Create the ground truths and detections scans of the targets;
Let’s start this example by creating the ground truths and the detections obtained by the two sensors.
We simulate the sensor detections by using a non-linear measurement model CartesianToBearingRange,
which allows to inverse-transform the detections to Cartesian coordinates. We simulate two
targets moving using ConstantVelocity transition model.
# instantiate the transition model
transition_model = CombinedLinearGaussianTransitionModel([ConstantVelocity(1e-4),
ConstantVelocity(1e-4)])
# Create a list of timesteps
timestamps = [start_time]
# create a set of truths
truths = set()
# Instantiate the groundtruth for the first target
truth = GroundTruthPath([GroundTruthState([0, 1, -100, 0.3], timestamp=start_time)])
# Loop over the simulation timesteps
for k in range(1, num_steps):
timeid = start_time + timedelta(seconds=scans_delay * k)
truth.append(GroundTruthState(
transition_model.function(truth[k-1], noise=True,
time_interval=timedelta(seconds=scans_delay)),
timestamp=timeid))
timestamps.append(timeid)
truths.add(truth)
# Add the second target, different starting position
truth = GroundTruthPath([GroundTruthState([0, 1, 100, -0.3], timestamp=start_time)])
# Loop over the simulation timesteps
for k in range(1, num_steps):
truth.append(GroundTruthState(
transition_model.function(truth[k-1], noise=True,
time_interval=timedelta(seconds=scans_delay)),
timestamp=start_time + timedelta(seconds=scans_delay*k)))
_ = truths.add(truth)
# Create the measurement model
from stonesoup.models.measurement.nonlinear import CartesianToBearingRange
sensor_1_mm = CartesianToBearingRange(
ndim_state=4,
mapping=(0, 2),
noise_covar=np.diag([np.radians(5), 10]))
# For simplicity let's assume both sensors have the same specifics
sensor_2_mm = deepcopy(sensor_1_mm)
# Collect the measurements using scans
scan_s1, scan_s2 = [], []
# Load the detections and clutter
from stonesoup.types.detection import TrueDetection
from stonesoup.types.detection import Clutter
from scipy.stats import uniform
# Loop over the steps
for k in range(num_steps):
detections1 = set()
detections2 = set()
# Introduce the delay every fifth scan
if not k%5 and k>0:
delay = extra_delay
else:
delay = 0
for truth in truths:
if np.random.rand() <= prob_detect: # for simplicity both scans get a detection at the same time
measurement = sensor_1_mm.function(truth[k], noise=True)
detections1.add(TrueDetection(state_vector=measurement,
groundtruth_path=truth,
timestamp=truth[k].timestamp))
measurement = sensor_2_mm.function(truth[k], noise=True)
detections2.add(TrueDetection(state_vector=measurement,
groundtruth_path=truth,
timestamp=truth[k].timestamp + timedelta(seconds=delay)))
# Generate clutter at this time-step
# randomly select a new x and y and calculate the Bearing-Range measurements
for _ in range(np.random.poisson(lambdaV)):
x1 = np.random.uniform(*v_bounds[0, :])
y1 = np.random.uniform(*v_bounds[1, :])
x2 = np.random.uniform(*v_bounds[0, :])
y2 = np.random.uniform(*v_bounds[1, :])
state1 = State(state_vector=StateVector([x1, truth.state_vector[1],
y1, truth.state_vector[3]]))
state2 = State(state_vector=StateVector([x2, truth.state_vector[1],
y2, truth.state_vector[3]]))
# Since the x-y locations are random, ignore the noise
detections1.add(Clutter(sensor_1_mm.function(state1, noise=False),
timestamp=truth[k].timestamp,
measurement_model=sensor_1_mm))
detections2.add(Clutter(sensor_2_mm.function(state2, noise=False),
timestamp=truth[k].timestamp + timedelta(seconds=delay),
measurement_model=sensor_2_mm))
scan_s1.append(detections1)
scan_s2.append(detections2)
2. Instantiate the tracking components;
We have a series of scans from the two sensors that contains true detections and some clutter.
The scans simulate the data obtained by the sensors, which contains both noise, in the form of clutter, and detections of the targets in the field of view.
It is time to instantiate the tracking components. For this example we consider
a UnscentedKalmanUpdater and UnscentedKalmanPredictor filter.
We use a probabilistic data associator using JPDA and PDAHypothesiser.
We consider a single updater since the measurement model is the same for
both sensors.
#Load the kalman filter components
from stonesoup.updater.kalman import UnscentedKalmanUpdater
from stonesoup.predictor.kalman import UnscentedKalmanPredictor
predictor = UnscentedKalmanPredictor(transition_model)
updater = UnscentedKalmanUpdater(measurement_model=sensor_1_mm)
# Load the data associator components
from stonesoup.dataassociator.probability import JPDA
from stonesoup.hypothesiser.probability import PDAHypothesiser
hypothesiser = PDAHypothesiser(
predictor=predictor,
updater=updater,
clutter_spatial_density=clutter_spatial_density,
prob_detect=prob_detect
)
# Set up the data associator
data_associator = JPDA(hypothesiser=hypothesiser)
# Instantiate the starting tracks
from stonesoup.types.track import Track
priori1 = GaussianState(state_vector=np.array([0, 1, -100, 0.3]),
covar=np.diag([1, 1, 1, 1]),
timestamp=start_time)
priori2 = GaussianState(state_vector=np.array([0, 1, 100, -0.3]),
covar=np.diag([1, 1, 1, 1]),
timestamp=start_time)
# prepare both the cases for using the algorithm C and excluding the OOSM
oosm_tracks = (Track([priori1]), Track([priori2]))
noOsm_tracks = (Track([priori1]), Track([priori2]))
3. Run the tracker and apply the algorithm on delayed measurements;
We can now run the tracker and generate the tracks.
By looping over the scans we can spot any OOS measurement and apply the algorithm. When we encounter a delayed measurement, at time \(\tau\), we make use of both the measurement model and transition model: first we use the measurement model inverse function at \(\tau\) to obtain a predicted Cartesian location of the target, then we use the transition model with time-inverse dynamics (\(t_{k} - \tau\)) to trace back where the target was in the scan-timestamp (\(t\)). With this pseudo-location, which is an approximation, we can compute the pseudo-measurement using the measurement model. In this way now we have a pseudo-measurement at \(t\) and we can keep the time-chain and process this as a true detection.
for k in range(len(scan_s1)): # loop over the scans
for detections_1, detections_2 in zip(scan_s1[k], scan_s2[k]):
# Check for the delay on the second scans
if timestamps[k] != detections_2.timestamp:
# Implementation of the algorithm C
# Use measurement-model inversion to obtain a predicted X-Y state
predicted_location = State(
state_vector=StateVector(sensor_2_mm.inverse_function(detections_2, noise=False)),
timestamp=timestamps[k])
# Use inverse dynamics to move from Tau-> t_K
oos_location = transition_model.function(predicted_location,
noise=False,
time_interval=detections_2.timestamp -
timestamps[k])
pseudo_state = State(state_vector=StateVector(oos_location),
timestamp=timestamps[k])
# Obtain the measurement detection/clutter from the pseudo-state
if type(detections_2) == TrueDetection:
pseudo_meas = TrueDetection(
state_vector=sensor_2_mm.function(pseudo_state, noise=False),
groundtruth_path=detections_2.groundtruth_path,
measurement_model=sensor_2_mm,
timestamp=pseudo_state.timestamp)
else:
pseudo_meas = Clutter(
state_vector=sensor_2_mm.function(pseudo_state, noise=False),
measurement_model=sensor_2_mm,
timestamp=pseudo_state.timestamp)
# Add the pseudo measurements
detections = [detections_1, pseudo_meas]
else:
# If the there is no delay
detections = [detections_1, detections_2]
# perform the association
associations = data_associator.associate(oosm_tracks, detections, timestamps[k])
# loop over the hypotheses
for track, hypotheses in associations.items():
for hypothesis in hypotheses:
if not hypothesis:
# if it is not an hypothesis, use the prediction
track.append(hypothesis.prediction)
else:
post = updater.update(hypothesis)
track.append(post)
Run the tracker while ignoring OOSM measurements
for k in range(len(scan_s1)):
for detections_1, detections_2 in zip(scan_s1[k], scan_s2[k]):
# Include only the In-sequence measurements
if timestamps[k] == detections_2.timestamp:
detections = [detections_1, detections_2]
associations = data_associator.associate(noOsm_tracks, detections, timestamps[k])
for track, hypotheses in associations.items():
for hyp in hypotheses:
if not hyp:
track.append(hyp.prediction)
else:
post = updater.update(hyp)
track.append(post)
4. Run the comparison between the resulting tracks and plot the results.
We have obtained the final tracks from the detections, and we have two sets of tracks, one with OOSM and one without. We can visualise the results and evaluate the track-to-track accuracy using the OSPA metric tool.
from stonesoup.plotter import AnimatedPlotterly
plotter = AnimatedPlotterly(timesteps=timestamps)
plotter.plot_ground_truths(truths, [0, 2])
plotter.plot_measurements(scan_s1, [0, 2], label='scan1', measurement_model=sensor_1_mm)
plotter.plot_measurements(scan_s2, [0, 2], label='scan2', measurement_model=sensor_1_mm)
plotter.plot_tracks(oosm_tracks, [0, 2], label='OOSM Tracks',
line= dict(color='orange'))
plotter.plot_tracks(noOsm_tracks, [0, 2], label='no-OOSM Tracks',
line= dict(color='red'))
plotter.fig
Evaluate the track-to-track accuracy
from stonesoup.metricgenerator.ospametric import OSPAMetric
ospa_oosm_truth = OSPAMetric(c=40, p=1, generator_name='OSPA_OOSM-truth',
tracks_key='oosm_track', truths_key='truths')
ospa_nooosm_truth = OSPAMetric(c=40, p=1, generator_name='OSPA_NoOOSM-truth',
tracks_key='noOosm_track', truths_key='truths')
from stonesoup.metricgenerator.manager import MultiManager
from stonesoup.dataassociator.tracktotrack import TrackToTruth
track_associator = TrackToTruth(association_threshold=30)
metric_manager = MultiManager([ospa_oosm_truth,
ospa_nooosm_truth], track_associator)
# add data to the metric manager
metric_manager.add_data({'truths': truths,
'oosm_track': oosm_tracks,
'noOosm_track': noOsm_tracks})
metrics = metric_manager.generate_metrics()
# load the metric plotter
from stonesoup.plotter import MetricPlotter
graph = MetricPlotter()
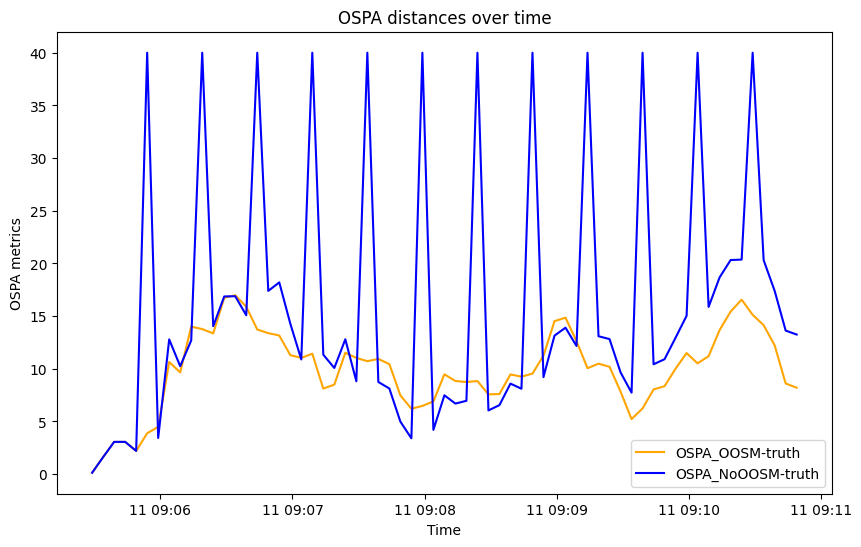
graph.plot_metrics(metrics, generator_names=['OSPA_OOSM-truth',
'OSPA_NoOOSM-truth'],
color=['orange', 'blue'])
graph.axes[0].set(ylabel='OSPA metrics', title='OSPA distances over time')
graph.fig
Conclusion
In this example we have explained how to deal with out of sequence detections using Algorithm C and time-inverse dynamics. In the OSPA metric summary presented we see that the difference between considering and ignoring OOSM can be significant, even in a simple scenario as this one considered.
References
Total running time of the script: (0 minutes 5.072 seconds)