Note
Click here to download the full example code or to run this example in your browser via Binder
11 - Gaussian mixture PHD tutorial
Background
Previous tutorials have described the difficulties of state estimation when there are multiple targets under consideration. The probability hypothesis density (PHD) filter has been proposed as a solution to this problem that is analogous to the Kalman Filter’s solution in single-object tracking. Where the Kalman filter propagates the first order movement of the posterior distribution of the target, the PHD filter creates a multiple target posterior distribution and propagates its first-order statistical moment, or PHD. At each time instance, the collections of targets and detections (including both measurements and false detections) are modelled as random finite sets. This means that the number of elements in each set is a random variable, and the elements themselves follow a probability distribution. Note that this is different from previously discussed filters which have a constant or known number of objects.
In a GM-PHD filter, each object is assumed to follow a linear Gaussian model, just like we saw in previous tutorials. However, the multiple objects need not have the same covariance matrices, meaning that the multiple target posterior distribution will be a Gaussian mixture (GM).
First we will recall some of the formulas that will be used in this filter.
Transition Density: Given a state \(p(\mathbf{x}_{k-1})\) at time \(k-1\), the probability density of a transition to the state \(p(\mathbf{x}_k)\) at time \(k\) is given by \(f_{k\vert k-1}(\mathbf{x}_{k}\vert \mathbf{x}_{k-1})\)
Likelihood Function: Given a state \(\mathbf{x}_{k}\) at time \(k\), the probability density of receiving the detection \(\mathbf{z}_{k}\) is given by \(g_{k}(\mathbf{z}_{k}\vert \mathbf{x}_{k})\)
The Posterior Density: The probability density of state \(\mathbf{x}_{k}\) given all the previous observations is denoted by \(p_{k}(\mathbf{x}_{k}\vert \mathbf{z}_{1:k})\). Using an initial density \(p_{0}(\cdot)\), we can apply Bayes’ recursion to show that the posterior density is actually
It is important to notice here that the state at time \(k\) can be derived wholly by the state at time \(k-1\).
Here we also introduce the following notation: \(p_{S,k}(\zeta)\) is the probability that a target \(S\) will exist at time \(k\) given that its previous state was \(\zeta\)
Suppose we have the random finite set \(\mathbf{X}_{k} \in \chi\) corresponding to the set of target states at time \(k\) and \(\mathbf{X}_k\) has probability distribution \(P\). Integrating over every region \(S \in \chi\), we get a formula for the first order moment (also called the intensity) at time \(k\), \(v_{k}\)
The set of targets spawned at time \(k\) by a target whose previous state was \(\zeta\) is the random finite set \(\mathbf{B}_{k|k-1}\). This new set of targets has intensity denoted \(\beta_{k|k-1}\).
The intensity of the random finite set of births at time \(k\) is given by \(\gamma_{k}\).
The intensity of the random finite set of clutter at time \(k\) is given by \(\kappa_{k}\).
The probability that a state \(x\) will be detected at time \(k\) is given by \(p_{D, k}(x)\).
Assumptions
The GM-PHD filter assumes that each target is independent of one another in both generated observations and in evolution. Clutter is also assumed to be independent of the target measurements. Finally, we assume that the target locations at a given time step are dependent on the multi-target prior density, and their distributions are Poisson. Typically, the target locations are also dependent on previous measurements, but that has been omitted in current GM-PHD algorithms.
Posterior Propagation Formula
Under the above assumptions, Vo and Ma 1 proved that the posterior intensity can be propagated in time using the PHD recursion as follows: \(\eqalignno{v _{ k\vert k-1} (x) =&\, \int p_{S,k}(\zeta)f_{k\vert k-1} (x\vert \zeta)v_{k-1}(\zeta)d\zeta\cr & +\int \beta_{k\vert k-1} (x\vert \zeta)v_{k-1}(\zeta)d\zeta+\gamma _{k}(x) & \cr v_{k} (x) =&\, \left[ 1-p_{D,k}(x)\right]v_{k\vert k-1}(x)\cr & +\!\!\sum\limits _{z\in Z_{k}} \!{{ p_{D,k}(x)g_{k}(z\vert x)}v_{k\vert k-1}(x) \over { \kappa _{k}(z)\!+\!\int p_{D,k}(\xi)g_{k}(z\vert \xi)v_{k\vert k-1}(\xi)}} . \cr &&}\)
For more information about the specific formulas for linear and non-linear Gaussian models, please see Vo and Ma’s full paper.
A Ground-Based Multi-Target Simulation
This simulation will include several targets moving in different directions accross the 2D Cartesian plane. The start locations of each object are random. These start locations are called priors and are known to the filter, via the density \(p_{0}(\cdot)\) discussed above.
At each time step, new targets are created and some targets die according to defined probabilities.
We start with some imports as usual.
# Imports for plotting
from matplotlib import pyplot as plt
plt.rcParams['figure.figsize'] = (14, 12)
plt.style.use('seaborn-colorblind')
# Other general imports
import numpy as np
from datetime import datetime, timedelta
start_time = datetime.now()
Generate ground truth
At the end of the tutorial we will plot the Gaussian mixtures. The ground truth Gaussian mixtures are stored in this list where each index refers to an instance in time and holds all ground truths at that time step.
truths_by_time = []
# Create transition model
from stonesoup.models.transition.linear import CombinedLinearGaussianTransitionModel, ConstantVelocity
transition_model = CombinedLinearGaussianTransitionModel(
(ConstantVelocity(0.3), ConstantVelocity(0.3)))
from stonesoup.types.groundtruth import GroundTruthPath, GroundTruthState
start_time = datetime.now()
truths = set() # Truths across all time
current_truths = set() # Truths alive at current time
start_truths = set()
number_steps = 20
death_probability = 0.005
birth_probability = 0.2
# Initialize 3 truths. This can be changed to any number of truths you wish.
truths_by_time.append([])
for i in range(3):
x, y = initial_position = np.random.uniform(-30, 30, 2) # Range [-30, 30] for x and y
x_vel, y_vel = (np.random.rand(2))*2 - 1 # Range [-1, 1] for x and y velocity
state = GroundTruthState([x, x_vel, y, y_vel], timestamp=start_time)
truth = GroundTruthPath([state])
current_truths.add(truth)
truths.add(truth)
start_truths.add(truth)
truths_by_time[0].append(state)
# Simulate the ground truth over time
for k in range(number_steps):
truths_by_time.append([])
# Death
for truth in current_truths.copy():
if np.random.rand() <= death_probability:
current_truths.remove(truth)
# Update truths
for truth in current_truths:
updated_state = GroundTruthState(
transition_model.function(truth[-1], noise=True, time_interval=timedelta(seconds=1)),
timestamp=start_time + timedelta(seconds=k))
truth.append(updated_state)
truths_by_time[k].append(updated_state)
# Birth
for _ in range(np.random.poisson(birth_probability)):
x, y = initial_position = np.random.rand(2) * [120, 120] # Range [0, 20] for x and y
x_vel, y_vel = (np.random.rand(2))*2 - 1 # Range [-1, 1] for x and y velocity
state = GroundTruthState([x, x_vel, y, y_vel], timestamp=start_time + timedelta(seconds=k))
# Add to truth set for current and for all timestamps
truth = GroundTruthPath([state])
current_truths.add(truth)
truths.add(truth)
truths_by_time[k].append(state)
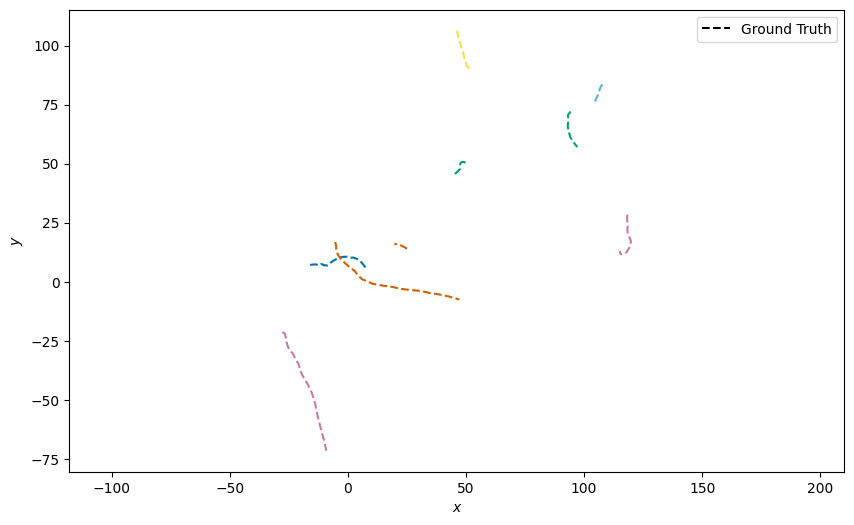
Plot the ground truth
from stonesoup.plotter import Plotter
plotter = Plotter()
plotter.plot_ground_truths(truths, [0, 2])
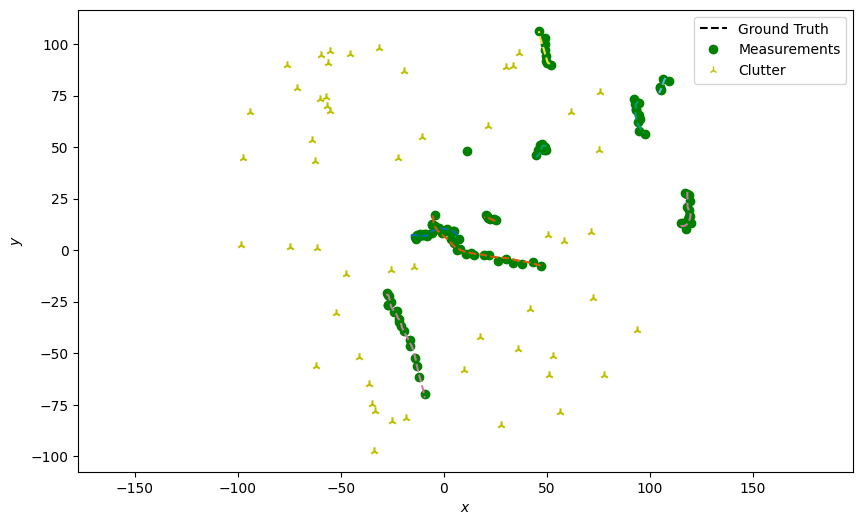
Generate detections with clutter
Next, generate detections with clutter just as in the previous tutorial. The clutter is assumed to be uniformly distributed accross the entire field of view, here assumed to be the space where \(x \in [-100, 100]\) and \(y \in [-100, 100]\).
# Make the measurement model
from stonesoup.models.measurement.linear import LinearGaussian
measurement_model = LinearGaussian(
ndim_state=4,
mapping=(0, 2),
noise_covar=np.array([[0.75, 0],
[0, 0.75]])
)
# Generate detections and clutter
from scipy.stats import uniform
from stonesoup.types.detection import TrueDetection
from stonesoup.types.detection import Clutter
all_measurements = []
# The probability detection and clutter rate play key roles in the posterior intensity.
# They can be changed to see their effect.
probability_detection = 0.9
clutter_rate = 3.0
for k in range(number_steps):
measurement_set = set()
timestamp = start_time + timedelta(seconds=k)
for truth in truths:
try:
truth_state = truth[timestamp]
except IndexError:
# This truth not alive at this time. Skip this iteration of the for loop.
continue
# Generate actual detection from the state with a 10% chance that no detection is received.
if np.random.rand() <= probability_detection:
# Generate actual detection from the state
measurement = measurement_model.function(truth_state, noise=True)
measurement_set.add(TrueDetection(state_vector=measurement,
groundtruth_path=truth,
timestamp=truth_state.timestamp,
measurement_model=measurement_model))
# Generate clutter at this time-step
for _ in range(np.random.poisson(clutter_rate)):
x = uniform.rvs(-100, 200)
y = uniform.rvs(-100, 200)
measurement_set.add(Clutter(np.array([[x], [y]]), timestamp=timestamp,
measurement_model=measurement_model))
all_measurements.append(measurement_set)
# Plot true detections and clutter.
plotter.plot_measurements(all_measurements, [0, 2], color='g')
plotter.fig
Create the Predicter and Updater
The updater is a PHDUpdater, and since it uses the mixed Gaussian paths, it is a
GM-PHD updater. For each individual track we use a KalmanUpdater. Here we assume
that the measurement range and clutter spatial density are known to the filter. We
invite you to change these variables to mimic removing this assumption.
from stonesoup.updater.kalman import KalmanUpdater
kalman_updater = KalmanUpdater(measurement_model)
# Area in which we look for target. Note that if a target appears outside of this area the
# filter will not pick up on it.
meas_range = np.array([[-1, 1], [-1, 1]])*200
clutter_spatial_density = clutter_rate/np.prod(np.diff(meas_range))
from stonesoup.updater.pointprocess import PHDUpdater
updater = PHDUpdater(
kalman_updater,
clutter_spatial_density=clutter_spatial_density,
prob_detection=probability_detection,
prob_survival=1-death_probability)
The GM-PHD filter quantifies the predicted-measurement distance, just as in previous
tutorials. The metric for this is the Mahalanobis distance. We also require two
objects which together form the predictor and to generate hypotheses (or predictions)
based on the previous state. Recall that the GM-PHD propagates the first-order
statistical moment which is a Gaussian mixture. The predicted state at the next time
step is also a Gaussian mixture and can be determined solely by the propagated prior.
Determining this predicted Gaussian mixture is the job for the
GaussianMixtureHypothesiser class. We must also generate a prediction for each track
in the simulation, and so use the DistanceHypothesiser object as before.
from stonesoup.predictor.kalman import KalmanPredictor
kalman_predictor = KalmanPredictor(transition_model)
from stonesoup.hypothesiser.distance import DistanceHypothesiser
from stonesoup.measures import Mahalanobis
base_hypothesiser = DistanceHypothesiser(kalman_predictor, kalman_updater, Mahalanobis(), missed_distance=3)
from stonesoup.hypothesiser.gaussianmixture import GaussianMixtureHypothesiser
hypothesiser = GaussianMixtureHypothesiser(base_hypothesiser, order_by_detection=True)
The updater takes a list of hypotheses from the hypothesiser and transforms them into
potential new states for our tracks. Each state is a TaggedWeightedGaussianState
object and has a state vector, covariance, weight, tag, and timestamp. Some of the
updated states have a very low weight, indicating that they do not contribute much to
the Gaussian mixture. To ease the computational complexity, a GaussianMixtureReducer
is used to merge and prune many of the states based on provided thresholds. States whose
distance is less than the merging threshold will be combined, and states whose weight
is less than the pruning threshold will be removed.
from stonesoup.mixturereducer.gaussianmixture import GaussianMixtureReducer
# Initialise a Gaussian Mixture reducer
merge_threshold = 5
prune_threshold = 1E-8
reducer = GaussianMixtureReducer(
prune_threshold=prune_threshold,
pruning=True,
merge_threshold=merge_threshold,
merging=True)
Now we initialize the Gaussian mixture at time k=0. In this implementation, the GM-PHD tracker knows the start state of the first 3 tracks that were created. After that it must pick up on new tracks and discard old ones. It is not necessary to provide the tracker with these start states, you can simply define the tracks as an empty set.
Feel free to change the state_vector from the actual truth state vector to something else. This would mimic if the tracker was unsure about where the objects were originating.
from stonesoup.types.state import TaggedWeightedGaussianState
from stonesoup.types.track import Track
from stonesoup.types.array import CovarianceMatrix
covar = CovarianceMatrix(np.diag([10, 5, 10, 5]))
tracks = set()
for truth in start_truths:
new_track = TaggedWeightedGaussianState(
state_vector=truth.state_vector,
covar=covar**2,
weight=0.25,
tag='birth',
timestamp=start_time)
tracks.add(Track(new_track))
The hypothesier takes the current Gaussian mixture as a parameter. Here we will initialize it to use later.
reduced_states = set([track[-1] for track in tracks])
To ensure that new targets get represented in the filter, we must add a birth component to the Gaussian mixture at every time step. The birth component’s mean and covariance must create a distribution that covers the entire state space, and its weight must be equal to the expected number of births per timestep. For more information about the birth component, see the algorithm provided in 2. If the state space is very large, it becomes inefficient to hold a component that covers it. Alternative implementations (as well as more dicussion about the birth component) are discussed in 3.
birth_covar = CovarianceMatrix(np.diag([1000, 2, 1000, 2]))
birth_component = TaggedWeightedGaussianState(
state_vector=[0, 0, 0, 0],
covar=birth_covar**2,
weight=0.25,
tag='birth',
timestamp=start_time
)
Run the Tracker
Now that we have all of our components, we can create a tracker. At each time instance, the tracker will go through four steps: hypothesise, update, reduce, and match. Let us briefly recap these four steps. The ‘hypothesise’ step is similar to the ‘prediction’ step in other filters. It uses the existing state space and the measurements to generate a list of all hypotheses for this time step (remember, each hypothesis is a Gaussian component). In the ‘update’ step, the filter combines the hypotheses into an updated Gaussian mixture. The ‘reduce’ step helps limit the computational complexity by merging and pruning the updated Gaussian mixture. The filter returns this final set of states and then we perform a ‘match’ step where we use the states’ tags to match them with an existing track (or create a new track).
These lists will be used to plot the Gaussian mixtures later. They are not used in the filter itself.
all_gaussians = []
tracks_by_time = []
We need a threshold to compare state weights against. If the state has a high enough weight in the Gaussian mixture, we will add it to an existing track or make a new track for it. Lowering this value makes the filter more sensitive but may also increase the number of false estimations. Increasing this value may increase the number of missed targets.
state_threshold = 0.25
for n, measurements in enumerate(all_measurements):
tracks_by_time.append([])
all_gaussians.append([])
# The hypothesiser takes in the current state of the Gaussian mixture. This is equal to the list of
# reduced states from the previous iteration. If this is the first iteration, then we use the priors
# defined above.
current_state = reduced_states
# At every time step we must add the birth component to the current state
if measurements:
time = list(measurements)[0].timestamp
else:
time = start_time + timedelta(seconds=n)
birth_component.timestamp = time
current_state.add(birth_component)
# Generate the set of hypotheses
hypotheses = hypothesiser.hypothesise(current_state,
measurements,
timestamp=time,
# keep our hypotheses ordered by detection, not by track
order_by_detection=True)
# Turn the hypotheses into a GaussianMixture object holding a list of states
updated_states = updater.update(hypotheses)
# Prune and merge the updated states into a list of reduced states
reduced_states = set(reducer.reduce(updated_states))
# Add the reduced states to the track list. Each reduced state has a unique tag. If this tag matches the tag of a
# state from a live track, we add the state to that track. Otherwise, we generate a new track if the reduced
# state's weight is high enough (i.e. we are sufficiently certain that it is a new track).
for reduced_state in reduced_states:
# Add the reduced state to the list of Gaussians that we will plot later. Have a low threshold to eliminate some
# clutter that would make the graph busy and hard to understand
if reduced_state.weight > 0.05: all_gaussians[n].append(reduced_state)
tag = reduced_state.tag
# Here we check to see if the state has a sufficiently high weight to consider being added.
if reduced_state.weight > state_threshold:
# Check if the reduced state belongs to a live track
for track in tracks:
track_tags = [state.tag for state in track.states]
if tag in track_tags:
track.append(reduced_state)
tracks_by_time[n].append(reduced_state)
break
else: # Execute if no "break" is hit; i.e. no track with matching tag
# Make a new track out of the reduced state
new_track = Track(reduced_state)
tracks.add(new_track)
tracks_by_time[n].append(reduced_state)
Plot the Tracks
First, determine the x and y range for axes. We want to zoom in as much as possible on the measurements and tracks while not losing any of the information. This section is not strictly necessary as we already set the field of view to be the range [-100, 100] for both x and y. However, sometimes an object may leave the field of view. If you want to ignore objects that have left the field of view, comment out this section and define the variables x_min = y_min = -100 and x_max = y_max = 100.
x_min, x_max, y_min, y_max = 0, 0, 0, 0
# Get bounds from the tracks
for track in tracks:
for state in track:
x_min = min([state.state_vector[0], x_min])
x_max = max([state.state_vector[0], x_max])
y_min = min([state.state_vector[2], y_min])
y_max = max([state.state_vector[2], y_max])
# Get bounds from measurements
for measurement_set in all_measurements:
for measurement in measurement_set:
if type(measurement) == TrueDetection:
x_min = min([measurement.state_vector[0], x_min])
x_max = max([measurement.state_vector[0], x_max])
y_min = min([measurement.state_vector[1], y_min])
y_max = max([measurement.state_vector[1], y_max])
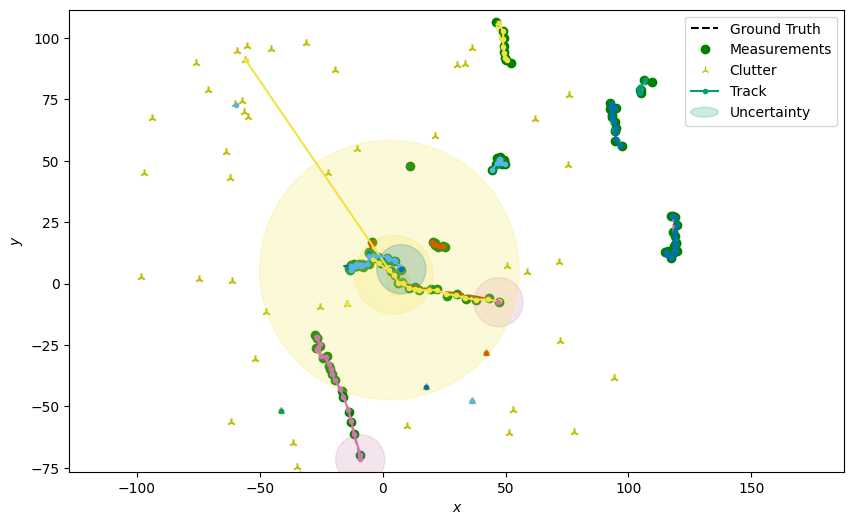
Now we can use the Plotter class to draw the tracks. Note that if the birth
component it plotted you will see its uncertainty ellipse centered around \((0, 0)\).
This ellipse need not cover the entire state space, as long as the distribution does.
# Plot the tracks
plotter = Plotter()
plotter.plot_ground_truths(truths, [0, 2])
plotter.plot_measurements(all_measurements, [0, 2], color='g')
plotter.plot_tracks(tracks, [0, 2], uncertainty=True)
plotter.ax.set_xlim(x_min-5, x_max+5)
plotter.ax.set_ylim(y_min-5, y_max+5)
Out:
(-76.55908760793943, 111.5015995926991)
Examining the Gaussian Mixtures
At every time step, the above GM-PHD algorithm creates a Gaussian mixture, which is a distribution over our target space. The following sections take a closer look at what this Gaussian really looks like. Note that the figures below only include the reduced states which have weight greater than 0.05. This decreases the overall number of states shown in the mixture and makes it easier to examine. But you can change this threshold parameter in the Run The Tracker section.
First we define a function that will help generate the z values for the Gaussian mixture. This lets us plot it later. This function has been updated from the one found here from this GPL-3.0 licensed repository.
from scipy.stats import multivariate_normal
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
def get_mixture_density(x, y, weights, means, sigmas):
# We use the quantiles as a parameter in the multivariate_normal function. We don't need to pass in any quantiles,
# but the last axis must have the components x and y
quantiles = np.empty(x.shape + (2,)) # if x.shape is (m,n) then quantiles.shape is (m,n,2)
quantiles[:, :, 0] = x
quantiles[:, :, 1] = y
# Go through each gaussian in the list and add its PDF to the mixture
z = np.zeros(x.shape)
for gaussian in range(len(weights)):
z += weights[gaussian]*multivariate_normal.pdf(x=quantiles, mean=means[gaussian, :], cov=sigmas[gaussian])
return z
For each timestep, create a new figure with 2 subplots. The plot on the left will show the 3D Gaussian Mixture density. The plot on the right will show a 2D birds-eye-view of the space, including the ground truths, detections, clutter, and tracks.
from matplotlib import animation
from matplotlib import pyplot as plt
from matplotlib.lines import Line2D # Will be used when making the legend
# This is the function that updates the figure we will be animating. As parameters we must
# pass in the elements that will be changed, as well as the index i
def animate(i, sf, truths, tracks, measurements, clutter):
# Set up the axes
axL.clear()
axR.set_title('Tracking Space at k='+str(i))
axL.set_xlabel("x")
axL.set_ylabel("y")
axL.set_title('PDF of the Gaussian Mixture')
axL.view_init(elev=30, azim=-80)
axL.set_zlim(0, 0.3)
# Initialize the variables
weights = [] # weights of each Gaussian. This is analogous to the probability of its existence
means = [] # means of each Gaussian. This is equal to the x and y of its state vector
sigmas = [] # standard deviation of each Gaussian.
# Fill the lists of weights, means, and standard deviations
for state in all_gaussians[i]:
weights.append(state.weight)
means.append([state.state_vector[0], state.state_vector[2]])
sigmas.append([state.covar[0][0], state.covar[1][1]])
means = np.array(means)
sigmas = np.array(sigmas)
# Generate the z values over the space and plot on the left axis
zarray[:, :, i] = get_mixture_density(x, y, weights, means, sigmas)
sf = axL.plot_surface(x, y, zarray[:, :, i], cmap=cm.RdBu, linewidth=0, antialiased=False)
# Make lists to hold the new ground truths, tracks, detections, and clutter
new_truths, new_tracks, new_measurements, new_clutter = [], [], [], []
for truth in truths_by_time[i]:
new_truths.append([truth.state_vector[0], truth.state_vector[2]])
for state in tracks_by_time[i]:
new_tracks.append([state.state_vector[0], state.state_vector[2]])
for measurement in all_measurements[i]:
if isinstance(measurement, TrueDetection):
new_measurements.append([measurement.state_vector[0], measurement.state_vector[1]])
elif isinstance(measurement, Clutter):
new_clutter.append([measurement.state_vector[0], measurement.state_vector[1]])
# Plot the contents of these lists on the right axis
if new_truths:
truths.set_offsets(new_truths)
if new_tracks:
tracks.set_offsets(new_tracks)
if new_measurements:
measurements.set_offsets(new_measurements)
if new_clutter:
clutter.set_offsets(new_clutter)
# Create a legend. The use of Line2D is purely for the visual in the legend
data_types = [Line2D([0], [0], color='white', marker='o', markerfacecolor='blue', markersize=15,
label='Ground Truth'),
Line2D([0], [0], color='white', marker='o', markerfacecolor='orange', markersize=15,
label='Clutter'),
Line2D([0], [0], color='white', marker='o', markerfacecolor='green', markersize=15,
label='Detection'),
Line2D([0], [0], color='white', marker='o', markerfacecolor='red', markersize=15,
label='Track')]
axR.legend(handles=data_types, bbox_to_anchor=(1.0, 1), loc='upper left')
return sf, truths, tracks, measurements, clutter
# Set up the x, y, and z space for the 3D axis
xx = np.linspace(x_min-5, x_max+5, 100)
yy = np.linspace(y_min-5, y_max+5, 100)
x, y = np.meshgrid(xx, yy)
zarray = np.zeros((100, 100, number_steps))
# Create the matplotlib figure and axes. Here we will have two axes being animated in sync.
# `axL` will be the a 3D axis showing the Gaussian mixture
# `axR` will be be a 2D axis showing the ground truth, detections, and updated tracks at
# each time step.
fig = plt.figure(figsize=(16, 8))
axL = fig.add_subplot(121, projection='3d')
axR = fig.add_subplot(122)
axR.set_xlim(x_min-5, x_max+5)
axR.set_ylim(y_min-5, y_max+5)
# Add an initial surface to the left axis and scattered points on the right axis. Doing
# this now means that in the animate() function we only have to update these variables
sf = axL.plot_surface(x, y, zarray[:, :, 0], cmap=cm.RdBu, linewidth=0, antialiased=False)
truths = axR.scatter(x_min-10, y_min-10, c='blue', linewidth=6, zorder=0.5)
tracks = axR.scatter(x_min-10, y_min-10, c='red', linewidth=4, zorder=1)
measurements = axR.scatter(x_min-10, y_min-10, c='green', linewidth=4, zorder=0.5)
clutter = axR.scatter(x_min-10, y_min-10, c='orange', linewidth=4, zorder=0.5)
# Create and display the animation
from matplotlib import rc
anim = animation.FuncAnimation(fig, animate, frames=number_steps, interval=500,
fargs=(sf, truths, tracks, measurements, clutter), blit=False)
rc('animation', html='jshtml')
anim
References
- 1
B. Vo and W. Ma, “The Gaussian Mixture Probability Hypothesis Density Filter,” in IEEE Transactions on Signal Processing, vol. 54, no. 11, pp. 4091-4104, Nov. 2006, doi: 10.1109/TSP.2006.881190
- 2
D. E. Clark, K. Panta and B. Vo, “The GM-PHD Filter Multiple Target Tracker,” 2006 9th International Conference on Information Fusion, 2006, pp. 1-8, doi: 10.1109/ICIF.2006.301809
- 3
B. Ristic, D. Clark, B. Vo and B. Vo, “Adaptive Target Birth Intensity for PHD and CPHD Filters,” in IEEE Transactions on Aerospace and Electronic Systems, vol. 48, no. 2, pp. 1656-1668, Apr 2012, doi: 10.1109/TAES.2012.6178085
Total running time of the script: ( 0 minutes 33.225 seconds)